Estimate and display direct and indirect effects of mediators and moderator in path models
Find the direct and indirect effects of a predictor in path models of mediation and moderation. Bootstrap confidence intervals for the indirect effects. Mediation models are just extended regression models making explicit the effect of particular covariates in the model. Moderation is done by multiplication of the predictor variables. This function supplies basic mediation/moderation analyses for some of the classic problem types.
mediate(y, x, m, data, mod = NULL, n.obs = NULL, use = "pairwise", n.iter = 5000, alpha = 0.05, std = FALSE,plot=TRUE) mediate.diagram(medi,digits=2,ylim=c(3,7),xlim=c(-1,10),show.c=TRUE, main="Mediation model",...) moderate.diagram(medi,digits=2,ylim=c(2,8),main="Moderation model",...)
Arguments
| y | The dependent variable (or a formula suitable for a linear model) |
|---|---|
| x | One or more predictor variables |
| m | One (or more) mediating variables |
| data | A data frame holding the data or a correlation or covariance matrix. |
| mod | A moderating variable, if desired |
| n.obs | If the data are from a correlation or covariance matrix, how many observations were used. This will lead to simulated data for the bootstrap. |
| use | use="pairwise" is the default when finding correlations or covariances |
| n.iter | Number of bootstrap resamplings to conduct |
| alpha | Set the width of the confidence interval to be 1 - alpha |
| std | standardize the covariances to find the standardized betas |
| plot | Plot the resulting paths |
| digits | The number of digits to report in the mediate.diagram. |
| medi | The output from mediate may be imported into mediate.diagram |
| ylim | The limits for the y axis in the mediate and moderate diagram functions |
| xlim | The limits for the x axis. Make the minimum more negative if the x by x correlations do not fit. |
| show.c | If FALSE, do not draw the c lines, just the partialed (c') lines |
| main | The title for the mediate and moderate functions |
| ... | Additional graphical parameters to pass to mediate.diagram |
Details
When doing linear modeling, it is frequently convenient to estimate the direct effect of a predictor controlling for the indirect effect of a mediator. See Preacher and Hayes (2004) for a very thorough discussion of mediation. The mediate function will do some basic mediation and moderation models, with bootstrapped confidence intervals for the mediation/moderation effects.
Functionally, this is just regular linear regression and partial correlation with some different output.
In the case of two predictor variables, X and M, and a criterion variable Y, then the direct effect of X on Y, labeled with the path c, is said to be mediated by the effect of x on M (path a) and the effect of M on Y (path b). This partial effect (a b) is said to mediate the direct effect of X --c--> Y: X --a -> M --b--> Y with X --c'--> Y where c' = c - ab.
Testing the significance of the ab mediation effect is done through bootstrapping many random resamples (with replacement) of the data.
For moderation, the moderation effect of Z on the relationship between X -> Y is found by taking the (centered) product of X and Z and then adding this XZ term into the regression.
In the case of being provided just a correlation matrix, the bootstrapped values are based upon bootstrapping from data matching the original covariance/correlation matrix with the addition of normal errors. This allows us to test the mediation/moderation effect even if not given raw data.
The function has been tested against some of the basic cases and examples in Hayes (2013) and the associated data sets.
Unless there is a temporal component that allows one to directly distinguish causal paths (time does not reverse direction), interpreting mediation models is problematic. Some people find it useful to compare the differences between mediation models where the causal paths (arrows) are reversed. This is a mistake and should not be done (Thoemmes, 2015).
For fine tuning the size of the graphic output, xlim and ylim can be specified in the mediate.diagram function. Otherwise, the graphics produced by mediate and moderate use the default xlim and ylim values.
Value
The total direct effect of x on y (c)
The beta effects of x (c') and m (b) on y
The indirect effect of x through m on y (c-ab)
mean bootstrapped value of indirect effect
Standard deviation of bootstrapped values
The upper and lower confidence intervals based upon the quantiles of the bootstrapped distribution.
The bootstrapped values themselves.
The effect of x on m
The effect of m on y
The interaction of x and mod (if specified)
References
Hayes, Andrew F. (2013) Introduction to mediation, moderation, and conditional process analysis: A regression-based approach. Guilford Press.
Preacher, Kristopher J and Hayes, Andrew F (2004) SPSS and SAS procedures for estimating indirect effects in simple mediation models. Behavior Research Methods, Instruments, \& Computers 36, (4) 717-731.
Thoemmes, Felix (2015) Reversing arrows in mediation models does not distinguish plausible models. Basic and applied social psychology, 27: 226-234.
Data from Hayes (2013), Preacher and Hayes (2004), and from Kerchoff (1974)
Note
There are a number of other packages that do mediation analysis (e.g., sem and lavaan) and they are probably preferred. This function is supplied for the more basic cases, with 1..k y variables, 1..n x variables, and 1 ..j mediators. It will not do two step mediation.
See also
setCor and setCor.diagram
Examples
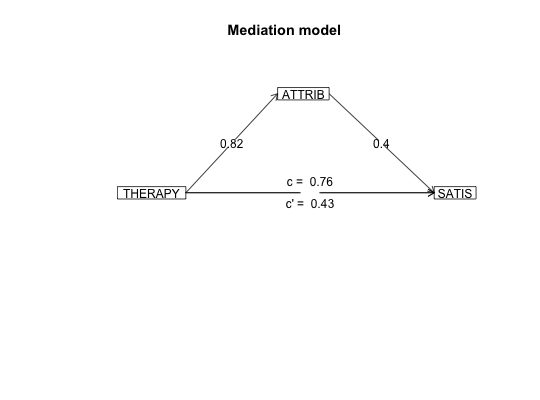
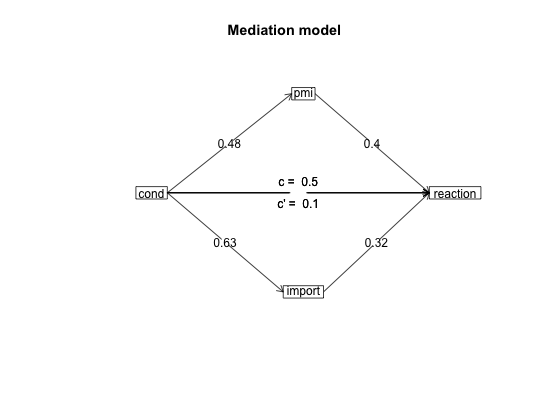
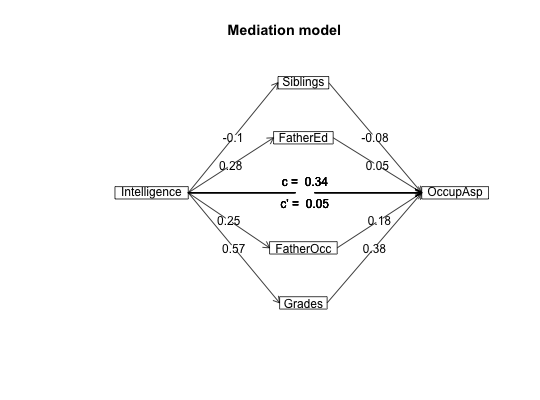
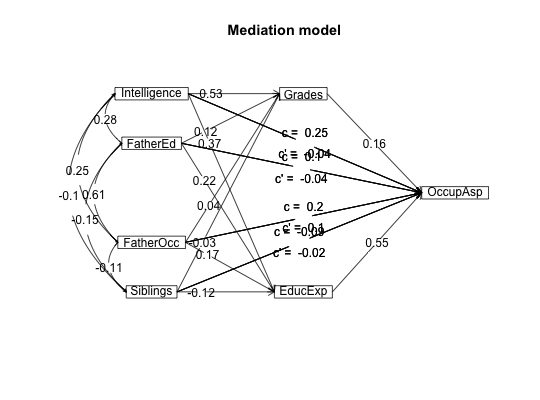
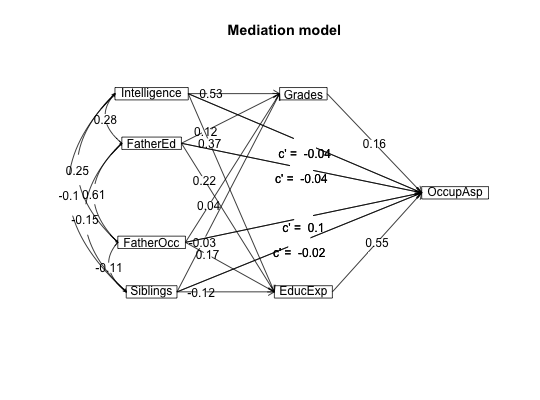
#data from Preacher and Hayes (2004) sobel <- structure(list(SATIS = c(-0.59, 1.3, 0.02, 0.01, 0.79, -0.35, -0.03, 1.75, -0.8, -1.2, -1.27, 0.7, -1.59, 0.68, -0.39, 1.33, -1.59, 1.34, 0.1, 0.05, 0.66, 0.56, 0.85, 0.88, 0.14, -0.72, 0.84, -1.13, -0.13, 0.2), THERAPY = structure(c(0, 1, 1, 0, 1, 1, 0, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 1, 0, 1, 0, 1, 1, 1, 0, 1, 1, 1, 1, 0), value.labels = structure(c(1, 0), .Names = c("cognitive", "standard"))), ATTRIB = c(-1.17, 0.04, 0.58, -0.23, 0.62, -0.26, -0.28, 0.52, 0.34, -0.09, -1.09, 1.05, -1.84, -0.95, 0.15, 0.07, -0.1, 2.35, 0.75, 0.49, 0.67, 1.21, 0.31, 1.97, -0.94, 0.11, -0.54, -0.23, 0.05, -1.07)), .Names = c("SATIS", "THERAPY", "ATTRIB" ), row.names = c(NA, -30L), class = "data.frame", variable.labels = structure(c("Satisfaction", "Therapy", "Attributional Positivity"), .Names = c("SATIS", "THERAPY", "ATTRIB"))) #n.iter set to 50 (instead of default of 5000) for speed of example mediate(1,2,3,sobel,n.iter=50) #The example in Preacher and Hayes#> Call: mediate(y = 1, x = 2, m = 3, data = sobel, n.iter = 50) #> #> The DV (Y) was SATIS . The IV (X) was THERAPY . The mediating variable(s) = ATTRIB . #> #> Total Direct effect(c) of THERAPY on SATIS = 0.76 S.E. = 0.31 t direct = 2.5 with probability = 0.019 #> Direct effect (c') of THERAPY on SATIS removing ATTRIB = 0.43 S.E. = 0.32 t direct = 1.35 with probability = 0.19 #> Indirect effect (ab) of THERAPY on SATIS through ATTRIB = 0.33 #> Mean bootstrapped indirect effect = 0.3 with standard error = 0.17 Lower CI = 0.02 Upper CI = 0.59 #> R2 of model = 0.31 #> To see the longer output, specify short = FALSE in the print statement #> #> Full output #> #> Total effect estimates (c) #> SATIS se t Prob #> THERAPY 0.76 0.31 2.5 0.0186 #> #> Direct effect estimates (c') #> SATIS se t Prob #> THERAPY 0.43 0.32 1.35 0.190 #> ATTRIB 0.40 0.18 2.23 0.034 #> #> 'a' effect estimates #> THERAPY se t Prob #> ATTRIB 0.82 0.3 2.74 0.0106 #> #> 'b' effect estimates #> SATIS se t Prob #> ATTRIB 0.4 0.18 2.23 0.034 #> #> 'ab' effect estimates #> SATIS boot sd lower upper #> THERAPY 0.33 0.3 0.17 0.02 0.59#the pmi covariance matrix from Hayes. 2013. #data set from Hayes, 2013 has 123 cases instead of the covariance matrix used here C.pmi <- structure(c(0.251232840197254, 0.119718779155005, 0.157470345195255, 0.124533519925363, 0.03052112488338, 0.0734039717446355, 0.119718779155005, 1.74573503931761, 0.647207783553245, 0.914575836332134, 0.0133613221378115, -0.0379181660669066, 0.157470345195255, 0.647207783553245, 3.01572704251633, 1.25128282020525, -0.0224576835932294, 0.73973743835799, 0.124533519925363, 0.914575836332134, 1.25128282020525, 2.40342196454751, -0.0106624017059843, -0.752990470478475, 0.03052112488338, 0.0133613221378115, -0.0224576835932294, -0.0106624017059843, 0.229241636678662, 0.884479541516727, 0.0734039717446355, -0.0379181660669066, 0.73973743835799, -0.752990470478475, 0.884479541516727, 33.6509729441557), .Dim = c(6L, 6L), .Dimnames = list(c("cond", "pmi", "import", "reaction", "gender", "age"), c("cond", "pmi", "import", "reaction", "gender", "age"))) #n.iter set to 50 (instead of default of 5000) for speed of example mediate(y="reaction",x = "cond",m=c("pmi","import"),data=C.pmi,n.obs=123,n.iter=50)#>#> Call: mediate(y = "reaction", x = "cond", m = c("pmi", "import"), data = C.pmi, #> n.obs = 123, n.iter = 50) #> #> The DV (Y) was reaction . The IV (X) was cond . The mediating variable(s) = pmi import . #> #> Total Direct effect(c) of cond on reaction = 0.5 S.E. = 0.28 t direct = 1.79 with probability = 0.077 #> Direct effect (c') of cond on reaction removing pmi import = 0.1 S.E. = 0.24 t direct = 0.43 with probability = 0.67 #> Indirect effect (ab) of cond on reaction through pmi import = 0.39 #> Mean bootstrapped indirect effect = 0.56 with standard error = 0.18 Lower CI = 0.19 Upper CI = 0.96 #> R2 of model = 0.33 #> To see the longer output, specify short = FALSE in the print statement #> #> Full output #> #> Total effect estimates (c) #> reaction se t Prob #> cond 0.5 0.28 1.79 0.0766 #> #> Direct effect estimates (c') #> reaction se t Prob #> cond 0.10 0.24 0.43 6.66e-01 #> pmi 0.40 0.09 4.26 4.04e-05 #> import 0.32 0.07 4.59 1.13e-05 #> #> 'a' effect estimates #> cond se t Prob #> pmi 0.48 0.24 2.02 0.0452 #> import 0.63 0.31 2.02 0.0452 #> #> 'b' effect estimates #> reaction se t Prob #> pmi 0.40 0.09 4.26 4.04e-05 #> import 0.32 0.07 4.59 1.13e-05 #> #> 'ab' effect estimates #> reaction boot sd lower upper #> cond 0.39 0.56 0.18 0.19 0.96#Data from sem package taken from Kerckhoff (and in turn, from Lisrel manual) R.kerch <- structure(list(Intelligence = c(1, -0.1, 0.277, 0.25, 0.572, 0.489, 0.335), Siblings = c(-0.1, 1, -0.152, -0.108, -0.105, -0.213, -0.153), FatherEd = c(0.277, -0.152, 1, 0.611, 0.294, 0.446, 0.303), FatherOcc = c(0.25, -0.108, 0.611, 1, 0.248, 0.41, 0.331), Grades = c(0.572, -0.105, 0.294, 0.248, 1, 0.597, 0.478 ), EducExp = c(0.489, -0.213, 0.446, 0.41, 0.597, 1, 0.651), OccupAsp = c(0.335, -0.153, 0.303, 0.331, 0.478, 0.651, 1 )), .Names = c("Intelligence", "Siblings", "FatherEd", "FatherOcc", "Grades", "EducExp", "OccupAsp"), class = "data.frame", row.names = c("Intelligence", "Siblings", "FatherEd", "FatherOcc", "Grades", "EducExp", "OccupAsp" )) #n.iter set to 50 (instead of default of 5000) for speed of demo mod.k <- mediate("OccupAsp","Intelligence",m= c(2:5),data=R.kerch,n.obs=767,n.iter=50)#>mediate.diagram(mod.k)#print the path values mod.k#> Call: mediate(y = "OccupAsp", x = "Intelligence", m = c(2:5), data = R.kerch, #> n.obs = 767, n.iter = 50) #> #> The DV (Y) was OccupAsp . The IV (X) was Intelligence . The mediating variable(s) = Siblings FatherEd FatherOcc Grades . #> #> Total Direct effect(c) of Intelligence on OccupAsp = 0.34 S.E. = 0.03 t direct = 9.83 with probability = 0 #> Direct effect (c') of Intelligence on OccupAsp removing Siblings FatherEd FatherOcc Grades = 0.05 S.E. = 0.04 t direct = 1.29 with probability = 0.2 #> Indirect effect (ab) of Intelligence on OccupAsp through Siblings FatherEd FatherOcc Grades = 0.29 #> Mean bootstrapped indirect effect = 0.27 with standard error = 0.03 Lower CI = 0.21 Upper CI = 0.31 #> R2 of model = 0.29 #> To see the longer output, specify short = FALSE in the print statement #> #> Full output #> #> Total effect estimates (c) #> OccupAsp se t Prob #> Intelligence 0.34 0.03 9.83 0 #> #> Direct effect estimates (c') #> OccupAsp se t Prob #> Intelligence 0.05 0.04 1.29 1.98e-01 #> Siblings -0.08 0.03 -2.59 9.91e-03 #> FatherEd 0.05 0.04 1.35 1.77e-01 #> FatherOcc 0.18 0.04 4.70 3.03e-06 #> Grades 0.38 0.04 10.03 0.00e+00 #> #> 'a' effect estimates #> Intelligence se t Prob #> Siblings -0.10 0.04 -2.78 0 #> FatherEd 0.28 0.03 7.97 0 #> FatherOcc 0.25 0.04 7.14 0 #> Grades 0.57 0.03 19.29 0 #> #> 'b' effect estimates #> OccupAsp se t Prob #> Siblings -0.08 0.03 -2.59 9.91e-03 #> FatherEd 0.05 0.04 1.35 1.77e-01 #> FatherOcc 0.18 0.04 4.70 3.03e-06 #> Grades 0.38 0.04 10.03 0.00e+00 #> #> 'ab' effect estimates #> OccupAsp boot sd lower upper #> Intelligence 0.29 0.27 0.03 0.21 0.31#Compare the following solution to the path coefficients found by the sem package mod.k2 <- mediate(y="OccupAsp",x=c("Intelligence","Siblings","FatherEd","FatherOcc"), m= c(5:6),data=R.kerch,n.obs=767,n.iter=50)#>mediate.diagram(mod.k2,show.c=FALSE) #simpler output#print the path values mod.k2#> Call: mediate(y = "OccupAsp", x = c("Intelligence", "Siblings", "FatherEd", #> "FatherOcc"), m = c(5:6), data = R.kerch, n.obs = 767, n.iter = 50) #> #> The DV (Y) was OccupAsp . The IV (X) was Intelligence Siblings FatherEd FatherOcc . The mediating variable(s) = Grades EducExp . #> #> Total Direct effect(c) of Intelligence on OccupAsp = 0.25 S.E. = 0.03 t direct = 7.29 with probability = 7.6e-13 #> Direct effect (c') of Intelligence on OccupAsp removing Grades EducExp = -0.04 S.E. = 0.03 t direct = -1.16 with probability = 0.25 #> Indirect effect (ab) of Intelligence on OccupAsp through Grades EducExp = 0.29 #> Mean bootstrapped indirect effect = 0.3 with standard error = 0.03 Lower CI = 0.25 Upper CI = 0.36 #> #> Total Direct effect(c) of Siblings on OccupAsp = -0.09 S.E. = 0.03 t direct = -2.78 with probability = 0.0056 #> Direct effect (c') of Siblings on NA removing Grades EducExp = -0.02 S.E. = 0.03 t direct = -0.68 with probability = 0.5 #> Indirect effect (ab) of Siblings on OccupAsp through Grades EducExp = -0.07 #> Mean bootstrapped indirect effect = 0.3 with standard error = 0.03 Lower CI = -0.09 Upper CI = -0.03 #> #> Total Direct effect(c) of FatherEd on OccupAsp = 0.1 S.E. = 0.04 t direct = 2.36 with probability = 0.018 #> Direct effect (c') of FatherEd on NA removing Grades EducExp = -0.04 S.E. = 0.04 t direct = -1.16 with probability = 0.25 #> Indirect effect (ab) of FatherEd on OccupAsp through Grades EducExp = 0.14 #> Mean bootstrapped indirect effect = 0.3 with standard error = 0.03 Lower CI = 0.13 Upper CI = 0.24 #> #> Total Direct effect(c) of FatherOcc on OccupAsp = 0.2 S.E. = 0.04 t direct = 4.8 with probability = 1.9e-06 #> Direct effect (c') of FatherOcc on NA removing Grades EducExp = 0.1 S.E. = 0.03 t direct = 2.85 with probability = 0.0044 #> Indirect effect (ab) of FatherOcc on OccupAsp through Grades EducExp = 0.1 #> Mean bootstrapped indirect effect = 0.3 with standard error = 0.03 Lower CI = 0.05 Upper CI = 0.15 #> R2 of model = 0.44 #> To see the longer output, specify short = FALSE in the print statement #> #> Full output #> #> Total effect estimates (c) #> OccupAsp se t Prob #> Intelligence 0.25 0.03 7.29 7.63e-13 #> Siblings -0.09 0.03 -2.78 5.60e-03 #> FatherEd 0.10 0.04 2.36 1.84e-02 #> FatherOcc 0.20 0.04 4.80 1.90e-06 #> #> Direct effect estimates (c') #> OccupAsp se t Prob #> Intelligence -0.04 0.03 -1.16 0.24600 #> Siblings -0.02 0.03 -0.68 0.49800 #> FatherEd -0.04 0.04 -1.16 0.24700 #> FatherOcc 0.10 0.03 2.85 0.00443 #> #> 'a' effect estimates #> Grades se t Prob #> Intelligence 0.53 0.03 17.16 0.00000 #> Siblings -0.03 0.03 -1.01 0.31300 #> FatherEd 0.12 0.04 3.16 0.00162 #> FatherOcc 0.04 0.04 1.09 0.27500 #> EducExp se t Prob #> Intelligence 0.37 0.03 12.45 0.00e+00 #> Siblings -0.12 0.03 -4.27 2.17e-05 #> FatherEd 0.22 0.04 6.00 2.99e-09 #> FatherOcc 0.17 0.04 4.63 4.28e-06 #> #> 'b' effect estimates #> OccupAsp se t Prob #> Grades 0.16 0.04 4.29 2.06e-05 #> EducExp 0.55 0.04 14.60 0.00e+00 #> #> 'ab' effect estimates #> OccupAsp boot sd lower upper #> Intelligence 0.29 0.30 0.03 0.05 0.15 #> Siblings -0.07 -0.06 0.01 0.05 0.15 #> FatherEd 0.14 0.18 0.03 0.05 0.15 #> FatherOcc 0.10 0.10 0.03 0.05 0.15